


Proteomics
Quantitative proteomics
At CSBMM, we will utilize high resolution mass spectrometry-based proteomics approaches to identify potential biomarkers in various cancers such as gastric cancer and head and neck cancer. Samples will be collected from various departments of Yenepoya Medical College and Yenepoya Dental College. Proteomic alterations in body fluids and tissues will be identified in precancerous lesions and cancerous tissues. The identified candidates may be utilized as early diagnostic markers in clinics in the future. Quantitative proteomic analysis will also be carried out for samples derived from infectious diseases.
Protein post-translational modifications and signaling networks
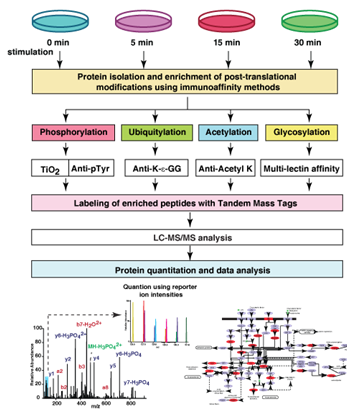
High resolution mass spectrometry-based proteomic methodologies will be developed at CSBMM. These strategies will be employed to monitor the dynamics of signaling complexes and unravel PTM cross talks including phosphorylation, ubiquitination, acetylation, sumoylation and nitrosylation in signaling pathways that are altered in various diseases with emphasis on tuberculosis, fungal infections and endemic febrile illnesses. The molecular networks thus identified will provide a platform for the development of novel biomarkers and therapeutic interventions, which will be essential for the better management of these diseases.
Targeted proteomics
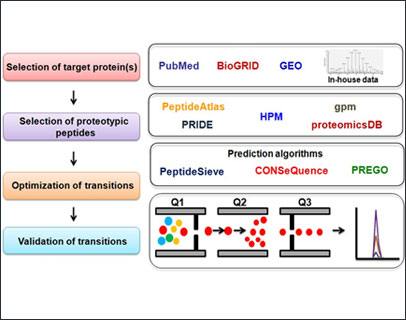
Targeted proteomic approaches such as multiple reaction monitoring (MRM) complement the widely used shotgun proteomic strategies. These techniques allow the accurate quantification of specific proteins of interest. Instruments such as QTRAP 6500 (SCIEX) and Orbitrap Fusion Tribrid (Thermo Scientific) mass spectrometers provide high resolving power, high-throughput ability that can be effectively used to analyze the abundance of specific proteins. At YU-IOB-CSBMM, we will be developing targeted proteomic assays to selectively identify pathogenic protein(s) and simultaneously quantitate their abundance in multiple clinical sample types. Similar workflows will be utilized to evaluate candidate biomarkers in various types of cancers.
Phytotherapeutics
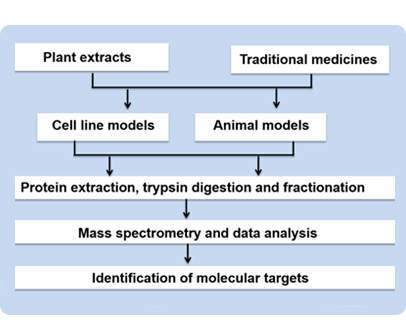
India has a great heritage in the use of traditional medicines dating back to several thousand years. The alternative forms of medicine include the extensive use of plants and herbs. These healthcare systems, although used to treat a plethora of diseases are critiqued for the lack of knowledge on their modes of action. At CSBMM, we will make use of proteomics and phosphoproteomic tools to gain insights into the molecular mechanisms of action of various plant extracts and traditional medicines. The knowledge gained therein may be used to develop alternate therapies using synthetic analogs against specific molecular targets. Further, as most medicinal plants indigenous to India remain unsequenced, genome information will be generated using Next-Gen Sequencing platforms to understand the pathways related to the synthesis of various secondary metabolites.
Plant proteomics
Due to sedentary mode of life, plants are frequently challenged by various harsh environmental conditions. During the process of evolution, plants have developed a set of versatile acclimation and adaptation strategies that help them withstand such harsh environmental fluctuations. Abiotic stresses such as salinity, extreme temperatures and water deficit limit the geographical locations and crop yields worldwide. Therefore, a better understanding of such responses is the key to decipher the regulatory mechanisms of better adaptation. At CSBMM, we will make use of gel-free proteomics and phosphoproteomics approach to decode the signaling networks involved in various abiotic and biotic stresses.
